Welcome
This web server provides online access to a series of tools developed by the Freiburg Bioinformatics Group. To start using it, please select from the listings below, or use the menu on the left. If you prefer doing a local installation on your machine, please visit our 'Download' section.
If you use our tools for research or education, please cite the corresponding articles from the 'Publications' section.
Version 5.0.12
MoDPepInt Server
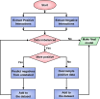
MoDPepInt (Modular Domain Peptide Interaction)
is a simple and interactive webserver, which
comprises three different tools, i.e. SH2PepInt, SH3PepInt and PDZPepInt,
for predicting the binding partners of three
different modular protein domains,
i.e. SH2, SH3 and PDZ domains, respectively.

SH2PepInt has been developed to predict binding partners to more than 50 human
SH2 domains. Depending on
the user requirement it uses PhosphoSitePlus and Gene Ontology
databases for predicting highly reliable SH2-peptide interactions.
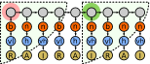
SH3PepInt is a fast and sophisticated graph kernel based tool
to predict SH3-peptide interactions. About 70 built-in models are
available for SH3-peptide predictions. It does not need any pre-alignment
of the peptides and uses Gene
Ontology database for getting more reliable interactions.
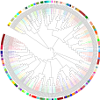
PDZPepInt is a cluster based prediction tool to predict binding
peptides of PDZ domains in human, mouse, fly and worm. About 40
built-in models that cover more than 220 PDZ domains across the species are
available. Gene Ontology
database can be used for getting reliable interactions. Additionally,
it will also consider the C-terminal peptides that are intrinsically
unstructured for getting high confidence interactions.
MoDPepInt is a meta-tool that runs interaction screens for
all domains-specific tools listed here. It might be used as a
starting point if no target domain information is known.
The Freiburg Bioinformatics Group also provides:
Freiburg RNA Tools
Freiburg RNA tools provides
online access to a series of RNA research tools developed by the
Freiburg Bioinformatics Group and colleagues for
sequence-structure alignments (LocARNA, CARNA, MARNA),
clustering (ExpaRNA),
interaction prediction (IntaRNA, CopraRNA, metaMIR),
identification of homologs (GLASSgo),
sequence design (AntaRNA, INFORNA, SECISDesign),
CRISPR repeat analyses (CRISPRmap),
and many more tasks.
CPSP-Tools Server
CPSP (Constraint-based Protein Structure Prediction)
is an exact and efficient approach to identify optimal structures
of lattice proteins within the hydrophobic-polar (HP) model. The
approach enables structure prediction within the 3D cubic and
3D face-centered-cubic (FCC) lattice for both backbone-only as well
as side-chain representing models.
Galaxy Project - Uni Freiburg
The Freiburg Galaxy Team offers a framework for scientists on
e.g. NGS data analyses (RNA-seq, ChIP-seq, Exome-seq, MethylC-seq),
genome annotation analyses for eukaryotic and prokaryotic organisms
(from gene prediction to functional description), Proteomics and
Metabolomics analysis, and the ChemicalToolBoX for analysis of
small compounds. Galaxy contains more than 800 different single
analysis tools and ready-to-use pipelines for different applications.
The Freiburg Galaxy Project is part of the “German Network for Bioinformatics Infrastructure” (Deutsches Netzwerk für Bioinformatik-Infrastruktur, de.NBI) and the Collaborative Research Centre (CRC) 992 for Medical Epigenetics and offers within the RNA Bioinformatic Centre (RBC) a central platform for RNA analysis.
The Freiburg Galaxy Project is part of the “German Network for Bioinformatics Infrastructure” (Deutsches Netzwerk für Bioinformatik-Infrastruktur, de.NBI) and the Collaborative Research Centre (CRC) 992 for Medical Epigenetics and offers within the RNA Bioinformatic Centre (RBC) a central platform for RNA analysis.